What is Phylogeny?
Phylogeny refers to the evolutionary relationships and relatedness between different species or populations. Phylogenetic trees can be constructed on the basis of phenotypic relationships or on genetic relationships[1].
Phylogeny of the PAH Protein
Once the homologs (and their sequences) are known, it is fairly easy to construct a phylogenetic tree. This involves inputting the sequences into a program like Clustal, which uses algorithms and statistics to predict evolutionary relationships between the homologs.
There are two different kinds trees that can be made: average distance or neighbor joining. Average distance trees use the alignments of homologous genes to infer closest relatives in context of the gene of interest. Neighbor joining trees, on the other hand, do not require alignments. Ultimately, neighbor joining trees are a form of an unrooted tree. That is, they show how many common ancestors are between two species, but does not infer how similar the sequences for a gene of interests are [2,3].
Additionally, each of these types of trees can be made using either % Identity or the BLOSUM62 matrix. Creating a tree with % Identity uses the actual sequences between homologs and how identical they are to create trees. BLOSUM62 references the likelihood of a certain amino acid being exchanged for another, and uses these likelihoods to determine how likely a certain mutation is bound to occur [4].
The problem with phylogeny lies in the fact that there are multiple algorithms that can be used. No algorithm is completely perfect, and as a result, depending on which algorithm is used, trees may look completely different from each other.
There are two different kinds trees that can be made: average distance or neighbor joining. Average distance trees use the alignments of homologous genes to infer closest relatives in context of the gene of interest. Neighbor joining trees, on the other hand, do not require alignments. Ultimately, neighbor joining trees are a form of an unrooted tree. That is, they show how many common ancestors are between two species, but does not infer how similar the sequences for a gene of interests are [2,3].
Additionally, each of these types of trees can be made using either % Identity or the BLOSUM62 matrix. Creating a tree with % Identity uses the actual sequences between homologs and how identical they are to create trees. BLOSUM62 references the likelihood of a certain amino acid being exchanged for another, and uses these likelihoods to determine how likely a certain mutation is bound to occur [4].
The problem with phylogeny lies in the fact that there are multiple algorithms that can be used. No algorithm is completely perfect, and as a result, depending on which algorithm is used, trees may look completely different from each other.
Discussion and Analysis
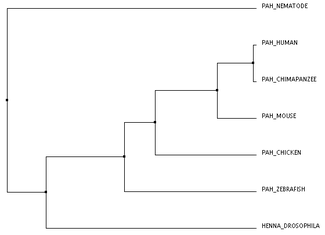
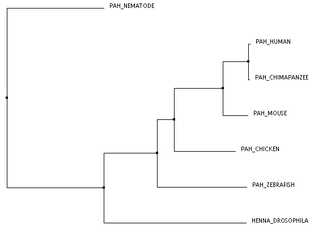
The trees above were created using Clustal's BLOSUM62 algorithm. Since BLOSUM62 is based upon statistics of amino acid substitution, it made since to use this algorithm while analyzing protein relatedness. Both of these trees show no discrepancies between them, and they both reflect the results from analyzing the homologous proteins with BLAST. That is, chimpanzees are most closely related to humans. This is followed by the mouse (another mammal), the chicken and zebrafish (other vertebrates), and finally by fruit flies and nematodes. Interestingly, nematodes are shown in these trees as a distinct outgroup. Using the BLOSUM62 algorithm for context, this shows that the nematodes' protein must have had some amino acid change(s) that wasn't expected or made it significantly different at a point in the protein than the other homologs.
References
1. "Learning with the ToL." Tree of Life: What Is Phylogeny. Tree of Life Project, 2004. Web. 17 Feb. 2013. <http://tolweb.org/tree/learn/concepts/whatisphylogeny.html>.
2. Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987 Jul;4(4):406-25. PubMed PMID: 3447015.
3. Opperdoes, Fred. "Neighbor-joining Method." Catholic University of Louvain, 8 Aug. 1997. Web. 18 Feb. 2013. <http://www.icp.ucl.ac.be/~opperd/private/neighbor.html>.
4. Staben, Chuck. "BLOSUM62 Matrix." BLOSUM62 Matrix. University of Kentucky, 28 Sept. 1998. Web. 17 Feb. 2013. <http://www.uky.edu/Classes/BIO/520/BIO520WWW/blosum62.htm>.
2. Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987 Jul;4(4):406-25. PubMed PMID: 3447015.
3. Opperdoes, Fred. "Neighbor-joining Method." Catholic University of Louvain, 8 Aug. 1997. Web. 18 Feb. 2013. <http://www.icp.ucl.ac.be/~opperd/private/neighbor.html>.
4. Staben, Chuck. "BLOSUM62 Matrix." BLOSUM62 Matrix. University of Kentucky, 28 Sept. 1998. Web. 17 Feb. 2013. <http://www.uky.edu/Classes/BIO/520/BIO520WWW/blosum62.htm>.