This page was produced as an assignment for Genetics 677, an undergraduate course at UW - Madison.
Final Thoughts
PKU is an interesting disease in the sense that the permanent and debilitating effects of an inherited disease can be offset by lifestyle choices. Though this is not particularly uncommon, I found PKU to be especially intriguing as many genetic disorders involving cognitive disabilities are only able to be dealt with and accepted instead of prevented. However, through reading testimonies and stories of people with PKU, prevention isn't always so simple. The formula that persons inflicted with PKU have to take in order to get protein is costly and often tastes so bad that patients have a hard time forcing themselves to take it. There have been foods that have been modified to contain no Phe, but they are extremely expensive. Sticking to the PKU diet requires such dedication and involves so much sacrifice that adults will frequently stray off diet, knowing that although having toxic levels of Phe will create a sort of "mental fog," their brains are developed to the point where it will no longer cause mental retardation. Furthermore, testing is not always accessible for those in underdeveloped countries or for those without access to hospitals and medical care. Given all these factors, PKU is also very much a socioeconomic disease as well.
Due to the prevalence of PKU (although relatively rare, it does affect thousands of people around the world) and the ability of the disease to be responsive to management, it should be made a priority to develop a more efficient way to manage PKU than the traditional diet. Medical professionals have known for a long time that this disease goes hand-in-hand with high levels of phenylalanine, that the diet works, etc., but there has been relatively little research done to provide concrete evidence on just how phenylalanine affects the body, the genome, and the proteome (or make-up of the proteins in the body) as a whole. There's very little incentive to find new solutions to the PKU problem when we already have one that works so well; if a baby is born with "X" level of phenylalanine, he/she should eat "X" amount of protein, and they'll be fine. However, this has mostly been found through trial and error, not scientific research, and it's possible that we're missing a lot of important information. It can't be particularly healthy to cut phenylalanine out of someone's diet unless it is absolutely critical, and now that genetic and proteomic knowledge has grown to the point that it has, we are in a better place to see just how different phenylalanine levels (low or high) can affect a body. Therefore, I propose a couple future research ideas that will help us better understand the effects of Phe levels on the body as a whole, not just on the areas which are known to be affected by PKU.
Due to the prevalence of PKU (although relatively rare, it does affect thousands of people around the world) and the ability of the disease to be responsive to management, it should be made a priority to develop a more efficient way to manage PKU than the traditional diet. Medical professionals have known for a long time that this disease goes hand-in-hand with high levels of phenylalanine, that the diet works, etc., but there has been relatively little research done to provide concrete evidence on just how phenylalanine affects the body, the genome, and the proteome (or make-up of the proteins in the body) as a whole. There's very little incentive to find new solutions to the PKU problem when we already have one that works so well; if a baby is born with "X" level of phenylalanine, he/she should eat "X" amount of protein, and they'll be fine. However, this has mostly been found through trial and error, not scientific research, and it's possible that we're missing a lot of important information. It can't be particularly healthy to cut phenylalanine out of someone's diet unless it is absolutely critical, and now that genetic and proteomic knowledge has grown to the point that it has, we are in a better place to see just how different phenylalanine levels (low or high) can affect a body. Therefore, I propose a couple future research ideas that will help us better understand the effects of Phe levels on the body as a whole, not just on the areas which are known to be affected by PKU.
Future Research Ideas
I have found, as a whole, that PKU is a clinically well-understood disease, and there has been a lot of valuable research done on the gene and protein itself. Biochemical mechanisms of the protein, inheritance patterns, types and frequencies of mutations, and physiological effects are all categorized and, in many ways, adequately described in the literature. Though there are specific studies about how PKU affects various functions (such as melanin production, cognitive ability, etc.), as stated above, I found a surprising lack of research into the whole-body effects that high phenylalanine can cause. Perhaps the full range of effects that PKU can have on the body can inferred and correctly guessed based off of theory alone, but, as shown by its ontology, phenylalanine metabolism affects a lot more than just its one specific pathway. This is why I propose the following question:
How do varying levels of Phe concentrations effect gene expression in various tissues?
One of the main concerns with PKU is the potential for severe and permanent mental retardation and brain damage. Because of this, much of the research on PKU has focused on this one area. A microarray (a technique used to measure what genes are being expressed in a specific tissue under specific conditions, such as starvation or exposure to a chemical) was performed on the brains mice with and without PKU to measure the effects of high Phe levels on gene expression in the brain [1]. However, I was able to find only minor amounts of information on how PKU and high Phe levels affects the rest of the body, especially in terms of gene expression.
One way to explore the effects of Phe levels in the body is to perform a series of microarrays. I propose controlling the blood Phe levels in mice (ranging from very low Phe to very high Phe, and with varying ages, from young to old) and performing microarrays on tissues such as the liver (due to its critical role in metabolism), the brain (although a couple microarrays have already been done, creating a range of Phe levels could help pinpoint what the concentration "critical point" for damage is), and the kidneys (as there has been indication that PAH expression is altered when certain kidney diseases are present). Knowing just how varying levels of Phe can affect them as a whole, individuals with PKU will be more equipped to make decisions as to how closely to stick to their diets.
To perform this experiment, blood Phe levels would be controlled in PKU-affected mice through diet of different ages (infancy, youth, maturity, etc.). Phe levels would range from 0 to up to 1000 micromole/L (a blood Phe level of around 100 is considered "normal"). A microarray analysis would be done from tissues taken from these mice to analyze what genes are being upregulated or downregulated due to the specific levels of phenylalanine.
Additionally, on a note that is related to the PAH gene and protein, but not so much related to PKU:
How is PAH connected to kidney disease (Chronic Renal Failure)?
Although PAH is known to be primarily housed in the liver, minor amounts of PAH expression has been detected in the kidneys as well [2]. It has been known for a long time that in the case of chronic renal failure (CRF), the amount of PAH protein expression in the kidneys is reduced even further [3,4]. This can result in toxic buildups of chemicals in the kidney (as is the case with PKU). Failure of PAH to function in the kidneys is a sign of advanced stage CRF. Indeed, though transcription of PAH in the kidneys seems to remain at its normal (although minimal) rate, activity of the protein is greatly reduced in those with CRF [3]. This suggests that the failure of PAH to express correctly in the kidneys is a result of translation or due to post-translational modifications. Since the PAH protein is known to have the ability to be activated partially through phosphorylation at the Serine residue in position 16 (for more information regarding PAH regulation, please visit the PAH Protein page), I predict that PAH in patients with CRF is being down-regulated by lack of phosphorylation at this site [5]. Another possibility is that the protein is being over-phosphorylated as a whole, and that extra phosphorylation in the catalytic domain is interrupting the catalytic activity.
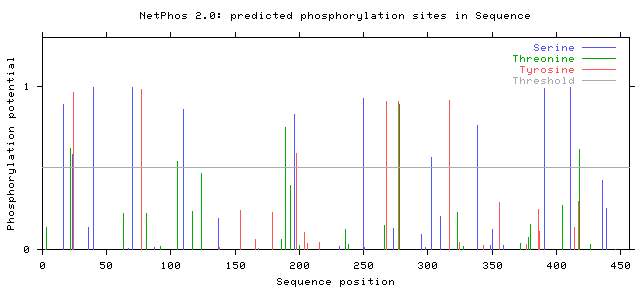
Using the NetPhos 2.0 server, I was able to predict the potential phosphorylation sites in the human PAH protein. This database turned up 22 sites (serines, threonines, and tyrosines) that had a significant chance of holding the ability to be phosphorylated. Of course, phosphorylation of any one of these sites does not necessarily guarantee or confer a unactivated protein. Many of these sites fall within the range of the catalytic domain.
How do varying levels of Phe concentrations effect gene expression in various tissues?
One of the main concerns with PKU is the potential for severe and permanent mental retardation and brain damage. Because of this, much of the research on PKU has focused on this one area. A microarray (a technique used to measure what genes are being expressed in a specific tissue under specific conditions, such as starvation or exposure to a chemical) was performed on the brains mice with and without PKU to measure the effects of high Phe levels on gene expression in the brain [1]. However, I was able to find only minor amounts of information on how PKU and high Phe levels affects the rest of the body, especially in terms of gene expression.
One way to explore the effects of Phe levels in the body is to perform a series of microarrays. I propose controlling the blood Phe levels in mice (ranging from very low Phe to very high Phe, and with varying ages, from young to old) and performing microarrays on tissues such as the liver (due to its critical role in metabolism), the brain (although a couple microarrays have already been done, creating a range of Phe levels could help pinpoint what the concentration "critical point" for damage is), and the kidneys (as there has been indication that PAH expression is altered when certain kidney diseases are present). Knowing just how varying levels of Phe can affect them as a whole, individuals with PKU will be more equipped to make decisions as to how closely to stick to their diets.
To perform this experiment, blood Phe levels would be controlled in PKU-affected mice through diet of different ages (infancy, youth, maturity, etc.). Phe levels would range from 0 to up to 1000 micromole/L (a blood Phe level of around 100 is considered "normal"). A microarray analysis would be done from tissues taken from these mice to analyze what genes are being upregulated or downregulated due to the specific levels of phenylalanine.
Additionally, on a note that is related to the PAH gene and protein, but not so much related to PKU:
How is PAH connected to kidney disease (Chronic Renal Failure)?
Although PAH is known to be primarily housed in the liver, minor amounts of PAH expression has been detected in the kidneys as well [2]. It has been known for a long time that in the case of chronic renal failure (CRF), the amount of PAH protein expression in the kidneys is reduced even further [3,4]. This can result in toxic buildups of chemicals in the kidney (as is the case with PKU). Failure of PAH to function in the kidneys is a sign of advanced stage CRF. Indeed, though transcription of PAH in the kidneys seems to remain at its normal (although minimal) rate, activity of the protein is greatly reduced in those with CRF [3]. This suggests that the failure of PAH to express correctly in the kidneys is a result of translation or due to post-translational modifications. Since the PAH protein is known to have the ability to be activated partially through phosphorylation at the Serine residue in position 16 (for more information regarding PAH regulation, please visit the PAH Protein page), I predict that PAH in patients with CRF is being down-regulated by lack of phosphorylation at this site [5]. Another possibility is that the protein is being over-phosphorylated as a whole, and that extra phosphorylation in the catalytic domain is interrupting the catalytic activity.
Using the NetPhos 2.0 server, I was able to predict the potential phosphorylation sites in the human PAH protein. This database turned up 22 sites (serines, threonines, and tyrosines) that had a significant chance of holding the ability to be phosphorylated. Of course, phosphorylation of any one of these sites does not necessarily guarantee or confer a unactivated protein. Many of these sites fall within the range of the catalytic domain.
In order to test if over-phosphorylation is potentially the problem for patients with CRF, I propose doing a proteomic analysis using immunoprecipitation and gel-based mass spectrometry. Using this technique, the current phosphorylation state of proteins can be analyzed. Samples from the kidneys of both healthy and CRF-inflicted mice will be collected. The PAH protein will be collected and purified using immunoprecipitation. Current phosphorylation states of the protein can be analyzed and compared using mass spectrometry.
This experiment can help give us insight as to why PAH is not functioning well in the kidney. If there is indeed under or over-phosphorylation, it is possible that this same phenomenon is happening to other important kidney proteins as well. This could provide some helpful information on patterns of malfunction in the end stages of Chronic Renal Failure.
Below you will find a copy of the presentation I gave about PKU:
This experiment can help give us insight as to why PAH is not functioning well in the kidney. If there is indeed under or over-phosphorylation, it is possible that this same phenomenon is happening to other important kidney proteins as well. This could provide some helpful information on patterns of malfunction in the end stages of Chronic Renal Failure.
Below you will find a copy of the presentation I gave about PKU:
| Final Presentation File | |
| File Size: | 10371 kb |
| File Type: | |
Resources
1. Surendran S, Matalon D, Tyring SK, Rady PL, Velagaleti GV, Matalon R. Altered expression of myocilin in the brain of a mouse model for phenylketonuria (PKU). Neurosci Lett. 2005 Jul 15;382(3):323-6. Epub 2005 Apr 2. PubMed PMID: 15925112.
2. Lichter-Konecki U, Hipke CM, Konecki DS. Human phenylalanine hydroxylase gene expression in kidney and other nonhepatic tissues. Mol Genet Metab. 1999 Aug;67(4):308-16. PubMed PMID: 10444341.
3. Wang M, Vyhmeister I, Swendseid ME, Kopple JD. Phenylalanine hydroxylase and tyrosine aminotransferase activities in chronically uremic rats. J Nutr. 1975;105:122–7.
4. Kopple JD. Phenylalanine and tyrosine metabolism in chronic kidney failure. J Nutr. 2007 Jun;137(6 Suppl 1):1586S-1590S; discussion 1597S-1598S. Review. PubMed PMID: 17513431.
5. Fitzpatrick PF. Allosteric regulation of phenylalanine hydroxylase. Arch Biochem Biophys. 2012 Mar 15;519(2):194-201. doi: 10.1016/j.abb.2011.09.012. Epub 2011 Oct 7. Review. PubMed PMID: 22005392; PubMed Central PMCID: PMC3271142.
6. Arntzen MØ, Osland CL, Raa CR, Kopperud R, Døskeland SO, Lewis AE, D'Santos CS. POSTMan (POST-translational modification analysis), a software application for PTM discovery. Proteomics. 2009 Mar;9(5):1400-6. doi: 10.1002/pmic.200800500. PubMed PMID: 19253288.
2. Lichter-Konecki U, Hipke CM, Konecki DS. Human phenylalanine hydroxylase gene expression in kidney and other nonhepatic tissues. Mol Genet Metab. 1999 Aug;67(4):308-16. PubMed PMID: 10444341.
3. Wang M, Vyhmeister I, Swendseid ME, Kopple JD. Phenylalanine hydroxylase and tyrosine aminotransferase activities in chronically uremic rats. J Nutr. 1975;105:122–7.
4. Kopple JD. Phenylalanine and tyrosine metabolism in chronic kidney failure. J Nutr. 2007 Jun;137(6 Suppl 1):1586S-1590S; discussion 1597S-1598S. Review. PubMed PMID: 17513431.
5. Fitzpatrick PF. Allosteric regulation of phenylalanine hydroxylase. Arch Biochem Biophys. 2012 Mar 15;519(2):194-201. doi: 10.1016/j.abb.2011.09.012. Epub 2011 Oct 7. Review. PubMed PMID: 22005392; PubMed Central PMCID: PMC3271142.
6. Arntzen MØ, Osland CL, Raa CR, Kopperud R, Døskeland SO, Lewis AE, D'Santos CS. POSTMan (POST-translational modification analysis), a software application for PTM discovery. Proteomics. 2009 Mar;9(5):1400-6. doi: 10.1002/pmic.200800500. PubMed PMID: 19253288.