This page was produced as an assignment for Genetics 677, an undergraduate course at UW - Madison.
What are Gene Motifs and Domains?
Gene motifs and domains help researchers to predict the function of the protein that a gene of interest may encode. Like protein domains, gene domains refer to the overall function of a protein the gene may encode, where as motifs refer to the actual sequence. In this sense, motifs are oftentimes more useful when comparing motifs and domains between species. Motifs are sections of highly conserved sequences that oftentimes have to do with transcription and/or translation (such as coding for a binding domain for transcription factors) of a gene of interest [1].
Motifs and Domains of the pah Gene
Motifs for the pah gene were found by submitting the human coding sequence (RNA) into the program MEME, which compares the sequence over databases for Human, Horse, Dog, and Mouse genome and motif sequences. It then generates a list of motifs that the gene most likely has. Submitting these motif sequences to GOMO will predict the function of these motifs [2,3].
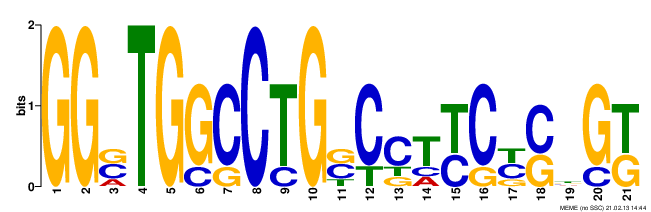
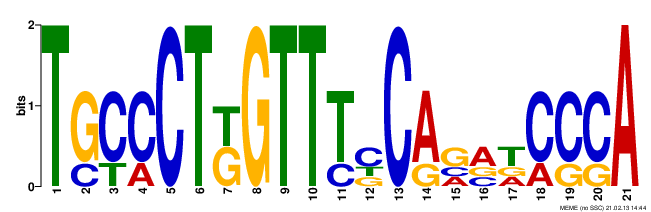
The images below can be a little difficult to interpret. Basically, the larger the nucleotide at a position is, the more common it was between the databases. For example, for Motif #1, the first two positions in the motif contained a "G" for all of the sequences that were compared. However, the third position in Motif #1 suggests that one of three nucleotides could be found between the sequences. Although there are a fair number of nucleotides that appear in the same position for all or most of the compared sequences, the amount of discrepancies between the sequences allows for many possibilities for potential functions of the motifs. Conveniently, GOMO lists the most likely potential functions and their statistical significance.
The images below can be a little difficult to interpret. Basically, the larger the nucleotide at a position is, the more common it was between the databases. For example, for Motif #1, the first two positions in the motif contained a "G" for all of the sequences that were compared. However, the third position in Motif #1 suggests that one of three nucleotides could be found between the sequences. Although there are a fair number of nucleotides that appear in the same position for all or most of the compared sequences, the amount of discrepancies between the sequences allows for many possibilities for potential functions of the motifs. Conveniently, GOMO lists the most likely potential functions and their statistical significance.
MEME and GOMO returned 3 motifs with predictable function for the pah gene. Because of the discrepancies and varieties in the sequences, there were large numbers (a few hundred) of predicted sequences for each motif. However, most of these predicted functions did not have a high identity or probability. As is common with DNA motifs, most of the predicted functions had to do with transcription and translation functions, and very little to do with the functions of the protein domains of the PAH protein.
References
1.Alberts, Bruce. "Gene Regulatory Proteins Were Discovered Using Bacterial Genetics." DNA-Binding Motifs in Gene Regulatory Proteins. U.S. National Library of Medicine, 18 Feb. 2002. Web. 28 Feb. 2013.
2.Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
3.Fabian A. Buske, Mikael Boden, Denis C. Bauer and Timothy L. Bailey, "Assigning roles to DNA regulatory motifs using comparative genomics", Bioinformatics, 26(7), 860-866, 2010.
2.Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
3.Fabian A. Buske, Mikael Boden, Denis C. Bauer and Timothy L. Bailey, "Assigning roles to DNA regulatory motifs using comparative genomics", Bioinformatics, 26(7), 860-866, 2010.