This page was produced as an assignment for Genetics 677, an undergraduate course at UW - Madison.
The Study of Homology
Protein homology refers to proteins in different species that have been inherited from a common ancestor [1]. Homologous proteins can also be considered orthologous if they have also retained the same function throughout time as well. Homology/orthology should not be confused with analogy, which refers to two proteins of the same function but evolved separately and independently from each other [1,2].
So why study homology? So what if a fly produces the same protein as a human? Well, this has a couple of implications. First of all, it is great news if a model organism (such as a fruit fly or a mouse) produces the same gene as a human, because it makes it possible - ethically, chronologically, and financially - to study that protein to a higher degree than would be possible to study in humans. Secondly, if a protein is conserved (that is, virtually unchanged, especially in function, throughout the course of evolution) in organisms from humans to fruit flies, it shows that the gene is probably involved in an important and/or necessary pathway or process [3]. A classic example of a highly conserved set of proteins are encoded by Hox genes, which help control development and body plans, and are present in animals from worms to humans and everything in between [4].
So why study homology? So what if a fly produces the same protein as a human? Well, this has a couple of implications. First of all, it is great news if a model organism (such as a fruit fly or a mouse) produces the same gene as a human, because it makes it possible - ethically, chronologically, and financially - to study that protein to a higher degree than would be possible to study in humans. Secondly, if a protein is conserved (that is, virtually unchanged, especially in function, throughout the course of evolution) in organisms from humans to fruit flies, it shows that the gene is probably involved in an important and/or necessary pathway or process [3]. A classic example of a highly conserved set of proteins are encoded by Hox genes, which help control development and body plans, and are present in animals from worms to humans and everything in between [4].
Finding the Homologs of the PAH Protein
Homologs were found using NCBI's BLAST (Basic Local Alignment Search Tool) program. The BLAST program takes inputs of sequences (such as mRNA, DNA, or protein sequences), compares them to a database of other sequences and genomes, and finds similar sequences. This program uses a set of algorithms and statistics to create alignments (areas of the sequence that match each other) and to predict how identical the sequences are overall. BLAST is a good starting place to find homologs - inputting a sequence of interest (such as the human PAH protein) and running it through the program will provide a list of potentially homologous genes in other species, based upon how identical the sequences are. Then, running the two potentially homologous protein against each other allows for analyzing alignments more finely. HomoloGene is another useful tool, which provides information on the known homologs of a specific protein of interest.
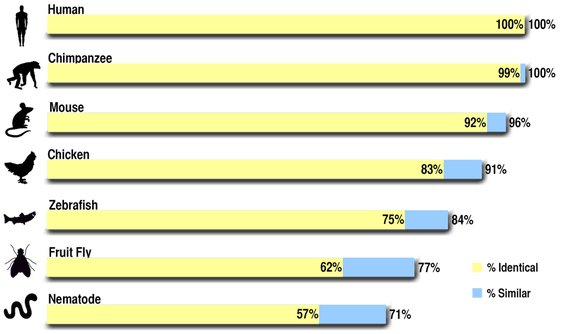
Comparing two proteins in BLAST provides two types of comparisons. Like when using the program to compare genes, it provides a list of homologs and how identical their sequences are (Max Identical). Additionally, it will provide information as to how similar the proteins are. This differs from the Max Identical score because many amino acids that make up proteins have similar chemical properties. In this sense, even though two proteins may not have identical sequences at a certain spot, those sequences could code for two very similar amino acids that would theoretically behave the same way chemically. Because of this flexibility, % Similar scores are higher than Max Identical Scores.
Comparing two proteins in BLAST provides two types of comparisons. Like when using the program to compare genes, it provides a list of homologs and how identical their sequences are (Max Identical). Additionally, it will provide information as to how similar the proteins are. This differs from the Max Identical score because many amino acids that make up proteins have similar chemical properties. In this sense, even though two proteins may not have identical sequences at a certain spot, those sequences could code for two very similar amino acids that would theoretically behave the same way chemically. Because of this flexibility, % Similar scores are higher than Max Identical Scores.
Discussion
Comparing the similarities between the genes, we can see that the human PAH protein is all but identical with its homolog in the chimpanzee, and is indeed 100% similar. This makes sense, as chimpanzees are humans' closest evolutionary relatives. The order of similarity (chimpanzee, mouse, chicken, zebrafish, fruit fly, and nematode) also can be considered intuitive, as all the mammals are more similar to the human PAH protein than the other animals, and all the vertebrates are more similar than the non-vertebrates. Additionally, the similarity between all of the proteins shows that PAH is conserved throughout the animal kingdom. The low E values (shown below) also show that these results are statistically significant, which allows inference that these proteins are all homologs and not analogs. There were no homologs to the PAH protein in non-animal organisms, such as E. coli, S. cerevisiae (yeast), and Arabidopsis.
PAH Protein Homologs References and Pages
|
Humans (Homo sapiens) - Phenylalanine Hydroxylase
Accession Number: NP_000268.1 GI Number: 4557819 FASTA Chimpanzee (Pan troglodytes) - Phenylalanine Hydroxylase
Accession Number: XP_001156919.1 GI Number: 114646575 FASTA E value: 0.0 Max identical: 99% % Similar: 100% Chicken (Gallus gallus) - Phenylalanine Hydroxylase
Accession Number: NP_001001298.1 GI Number: 47604920 FASTA E value: 0.0 Max identical: 83% % Similar: 91% Zebrafish (Danio rerio) - Phenylalanine Hydroxylase
Accession Number: NP_956845.1 GI Number: 41054599 FASTA E value: 0.0 Max identical: 75% % Similar: 84% |
Mouse (Mus musculus) - Phenylalanine Hydroxylase
Accession Number: NP_032803.2 GI Number: 171543886 FASTA E value: 0.0 Max identical: 92% % Similar: 96% Nematode (Caenorhabditis elegans) - Phenylalanine Hydroxylase
Accession Number: NP_001254184.1 GI Number: 392891006 FASTA E value: 0.0 Max identical: 57% % Similar: 71% Fruit Fly (Drosophila melanogaster) - Henna
Accession Number: NP_523963.2 GI Number: 24660393 FASTA E value: 0.0 Max identical: 62% % Similar: 77& |
| pah_protein_alignments.clustalw.webarchive | |
| File Size: | 5 kb |
| File Type: | webarchive |
References
1. Delsuc F, Brinkmann H, Philippe H. Phylogenomics and the reconstruction of the tree of life. Nat Rev Genet. 2005 May;6(5):361-75. Review. PubMed PMID: 15861208.
2. J. H. Jackson Laboratory. "Gene Similarity: Some Definitions." Michigan State University, 08 Apr. 1999. Web. <https://www.msu.edu/~jhjacksn/Reports/similarity.htm>.
3. Brody, Thomas B., PhD. "Evolutionarily Conserved Developmental Pathways." The Interactive Fly. Society for Developmental Biology, 10 Feb. 2012. Web. 15 Feb. 2013. <http://www.sdbonline.org/fly/aimain/aadevinx.htm>.
4. Lemons D, McGinnis W. Genomic evolution of Hox gene clusters. Science. 2006 Sep 29;313(5795):1918-22. Review. PubMed PMID: 17008523.
2. J. H. Jackson Laboratory. "Gene Similarity: Some Definitions." Michigan State University, 08 Apr. 1999. Web. <https://www.msu.edu/~jhjacksn/Reports/similarity.htm>.
3. Brody, Thomas B., PhD. "Evolutionarily Conserved Developmental Pathways." The Interactive Fly. Society for Developmental Biology, 10 Feb. 2012. Web. 15 Feb. 2013. <http://www.sdbonline.org/fly/aimain/aadevinx.htm>.
4. Lemons D, McGinnis W. Genomic evolution of Hox gene clusters. Science. 2006 Sep 29;313(5795):1918-22. Review. PubMed PMID: 17008523.